From our partners at AKOYA, latest insights on spatial imaging for intestines, breast and kidney tissue
24th Oct 2023
Bringing order to our inner cosmos: New spatial maps of the intestines, breast, and kidney reveal valuable insights
- Published: September 13, 2023
There are 37.2 trillion cells in the human body – 100 times more than the number of stars in the Milky Way. Making sense of the chaos of our inner cosmos is the aim of the Human Cell Atlas Project and Human BioMolecular Atlas Program (HuBMAP). This summer, three new studies from consortium members shed further light on the hidden order within our tissues — using multiplexed imaging, spatial biology, and other technologies to reveal valuable insights into our inner world of cellular communities. 2
Table of contents
Charting a course through the gut
In humans, the proper functioning of organs and tissues is dependent on the interaction, spatial organization, and specialization of all our cells. With so many cells to account for, determining each cell’s identity and function along with its relationship to neighboring cells is a monumental undertaking. So, scientists have started to harness a combination of cutting-edge single-cell techniques to create unique “ID cards” for each cell type (based on which of the 20,000 genes in that individual cell are switched on and which proteins are expressed). Then, applying advanced imaging and analysis techniques, they map these individual cells to precise locations in organs and tissues to understand their functions and relationships with their neighbors.
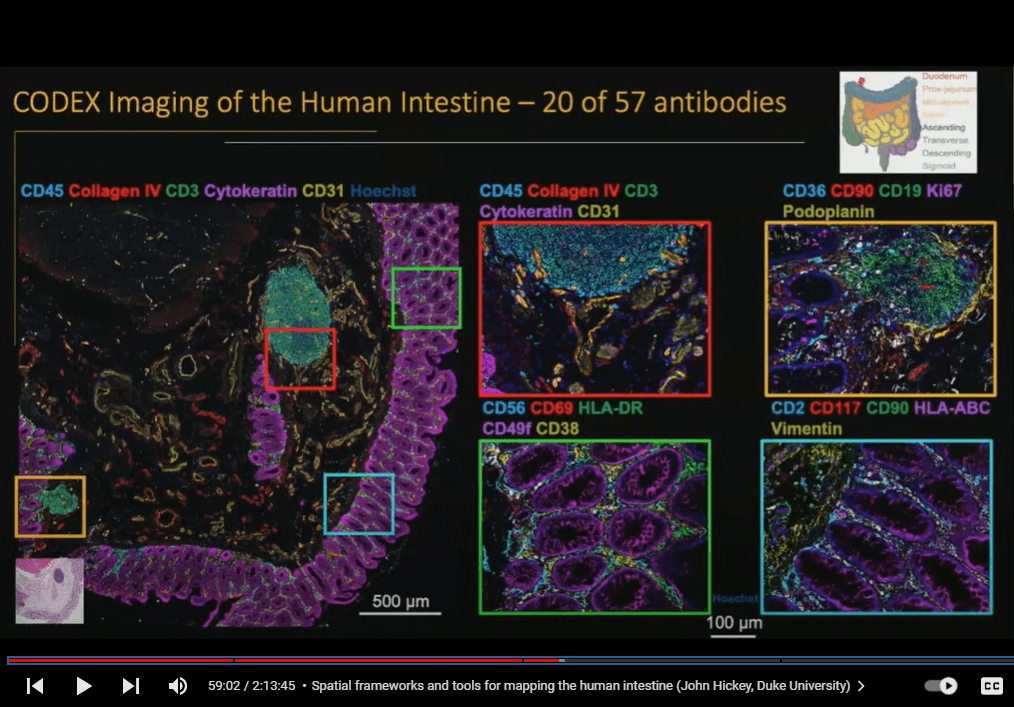
To map the intestine (which is more than 9 meters or 25 feet long), Stanford University scientists examined eight regions of the small and large intestine from nine donors using the PhenoCycler spatial phenotyping platform.
Published in Nature as “Organization of the human intestine at single-cell resolution,” the work highlighted how cell compositions differ substantially across regions of the intestine with many complex subtypes enacting different functions. These include epithelial cells that make up the intestinal lining, connective tissue cells, nerve cells, and immune cells. The PhenoCycler platform enabled scientists to pinpoint not only where each cell type is located, but also which other cells they associate with. They identified 20 distinct cellular neighborhoods, as well as distinct immunological niches.
“It was a bit like exploring a new planet, in that we didn’t know exactly what cell types we would find or how they would be organized,” Michael Snyder (@SnyderShot), professor and chair of genetics at Stanford School of Medicine, told Genetic Engineering and Biotechnology News, adding that “this will be foundational for our understanding of all kinds of digestive diseases.”
To learn more, check out this presentation by Dr. John Hickey.
The highways and byways of the breast
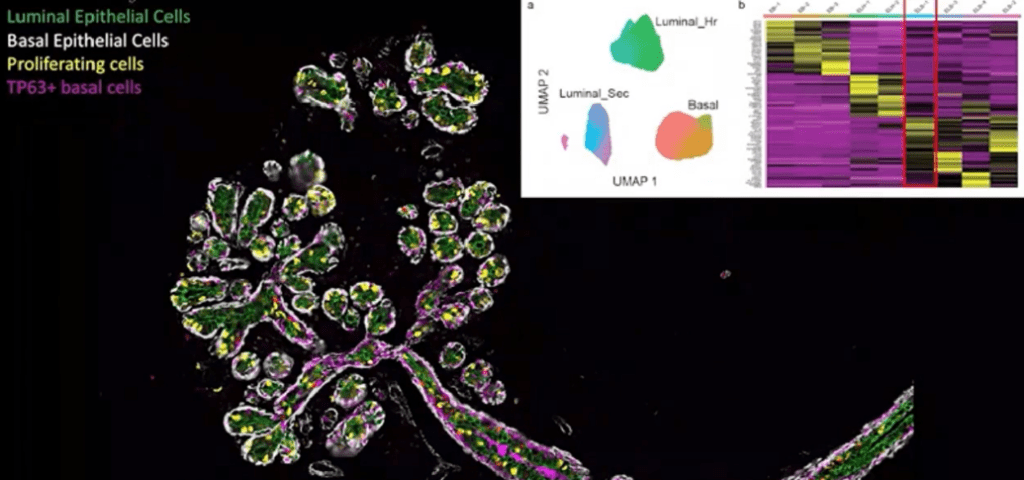
Just as in the intestine and other organs, cells in the breast are embedded into a complex three-dimensional microenvironment consisting of the extracellular matrix as well as other cell types. The breast epithelial system consists of a ductal epithelial network, with four distinct regions of interest: lobular units, connective tissue, ducts, and adipose tissue.
Using the PhenoCycler platform and the StarDist pipeline, a team led by Kai Kessenbrock (@KessenbrockLab), Assistant Professor of Biological Chemistry at UC Irvine, conducted a spatial phenotyping analysis of healthy breast tissue labeled with 34 different antibodies. This rich imaging data (containing more than 81,000 individual cells) was turned into a cell by protein matrix to study cell density and diversity.
They found that lobules have extremely high population density and diversity – these are the cities and metropolitan areas of the breast – while ducts corresponded to interstate highways, and connective and adipose tissue displayed the same low density and diversity expected in suburban and rural areas. They also performed cellular proximity analysis to characterize typical neighboring cells, identifying 14 distinct cellular neighborhoods.
Dr. Kessenbrock’s team extended their investigation into non-epithelial cell types as well. To-date, the most research has focused on epithelial cells layers as these undergo dramatic transformation during menstruation, pregnancy, and lactation. As a result, less is known about non-epithelial cells. The team profiled 535,941 cells from 62 women and 120,024 nuclei from 20 women, identifying 11 major cell types and 53 cell states. As described in their Nature paper, “A spatially resolved single-cell genomic atlas of the adult human breast”, their data revealed abundant pericyte, endothelial and immune cell populations, and highly diverse luminal epithelial cell states.
The multiomics approach allowed the team to develop a comprehensive map of the distinct cell populations, pathways, lineage hierarchies, and cellular neighborhoods in human breast tissue.
The spatial mapping, in particular, “revealed an unexpectedly rich ecosystem of tissue-resident immune cells in the ducts and lobules, as well as distinct molecular differences between ductal and lobular regions,” the team reported. As reported separately in this whitepaper, the PhenoCycler platform’s ability to provide single-cell resolution further helped identify several rare cell types that were missed by other technologies that use a regions of interest imaging approach.
You can check out this webinar with Dr. Kessenbrock to learn more.
Welcome to the renal neighborhood
If you’ve ever had a kidney stone, you may have wondered what exactly it was and how and why it got there. Despite decades of innovation and efforts by researchers to describe its pathophysiology, the precise mechanisms contributing to kidney stone formation remain poorly understood.
However, a team of researchers from the Indiana University School of Medicine have brought us several steps closer to an answer after defining the spatial molecular landscape and specific pathways contributing to stone-mediated injury in the human papilla and identifying associated urinary biomarkers.
In the recent study, published in Nature Communications as “A spatially anchored transcriptomic atlas of the human kidney papilla identifies significant immune injury in patients with stone disease”, the team described how they took a multiomics approach to create a first-of-its-kind spatial map of the kidney.
Leveraging a comprehensive single nuclear RNA sequencing (snRNAseq) profile (over 200,000 cells) of the adult human kidney from the Human Biomolecular Atlas (HubMAP) and the Kidney Precision Medicine project (KPMP) consortia as a foundation, the Indiana researchers delved into cell type diversity in the human kidney papillae (the areas where the openings of the collecting ducts enter the kidney and where urine flows into the ureters), with an eye on unique features compared to other kidney cell types, including the renal cortex (the outer layer of the kidney, responsible for filtering waste products from the blood) and the medulla (the innermost layer, where the nephrons are located, responsible for regulating electrolyte levels).
Applying ultrahigh-plex PhenoCycler analysis of reference tissues with 32 cell markers, the team not only validated the cellular diversity in the papilla, but also identified new cell populations with plausible biological significance.
A full landscape of papillary cells was defined, including the presence of papillary surface epithelial cells, stromal and immune cells, unique subtypes of principal cells, and an undifferentiated epithelial cell type that localized to regions of injury or mineral deposition.
They were also able to trace injury pathways, as well as immune system process/leukocyte (myeloid) immune activation, response to oxidative stress and extracellular matrix organization.
“Cumulatively, our data suggest stone disease alters the phenotype of collecting duct and other tubular cells toward an injury phenotype. These changes may trigger immune and fibrogenic responses that eventually replace the healthy tubular epithelial make-up of the papilla,” the authors wrote.
What the maps tell us
Without maps of different cell types, their molecular characteristics, and where they are located in the body, we cannot describe all their functions and understand the networks that direct their activities. The same immune cell placed in a different context within the tumor can play a completely different role, for instance.
Although the primary intention of the maps developed by the HCA and HuBMAP initiatives is to serve as a reference of healthy organs and how they operate and develop over time, they can also be used to compare to organs in different disease states.
The Stanford University team found it useful to investigate the organization of healthy tissue and how it changed throughout the digestive tract. They found that what’s normal in one region might be a sign of disease in another. They also found other associations. Donors with higher body mass index (BMI) had a greatly increased number of M1 macrophages (which are associated with inflammation) and those with a history of hypertension also had fewer CD8+ T cells, which play a role in seeking out and destroying possible cancer cells.
As the breast study noted, understanding the variation and diversity of the immune cells is very important for breast cancer, where immunotherapy has recently become the standard of care for some subtypes. And the team was able to integrate data that identified significant changes in the breast tissue architecture that corresponded to menopause, age, and BMI.
The kidney mapping exercise also led to a bonus finding: a potential biomarker. MMP7 and MMP9 were identified as two proteins linked to active stone disease and mineralization within the papilla, with levels of MMP9 and MMP7 in the urine significantly higher in patients with history of stone disease compared to healthy controls and correlated with disease activity.
We look forward to delving into more exciting science enabled by the PhenoCycler and PhenoImager platforms in future posts. To stay updated on our ongoing series celebrating 1,000+ publications featuring Akoya’s spatial biology technologies:
- Keep visiting our blog
- Follow #AkoyaInAction on LinkedIn
